Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose...
Fantastic news! We've Found the answer you've been seeking!
Question:

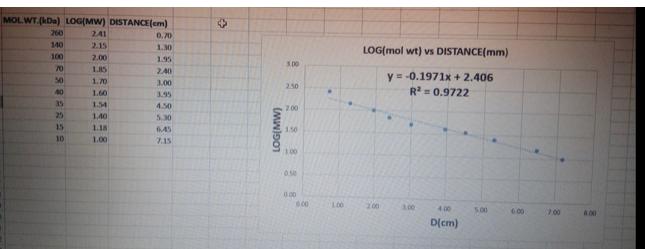

Transcribed Image Text:
Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve Using the migration of the DNA markers (Invitrogen 1 Kb Plus DNA ladder) on the agarose gel with your PCR product you should construct a standard curve of the migration rate of the DNA markers against the log10 value of their molecular weight. The table below is provided to tabulate the data necessary for making this graph. NOTE: Only the middle column (Log10 size of DNA marker) will be graded. Do not enter any units (i.e. bp) in this column and give your answer to 2 significant figures. Molecular weight and migration of DNA markers Size of DNA marker (bp) 3000 2000 1500 1000 850 650 500 400 300 200 100 Logo size of DNA marker (bp) Distance migrated by DNA marker from top of agarose gel (mm) MOLWT.(kDa) LOG(MW) DISTANCE(cm) 241 2.15 2.00 1.85 1.70 1.60 1.54 1,40 1.18 1.00 260 140 100 553588888 70 50 40 25 10 0.70 1.30 1.95 2.40 3.00 3.95 4.50 5.30 6.45 7.15 + (MW)901 5.00 2.50 200 1.50 1.00 0.500 0.00 0.00 100 LOG(mol wt) vs DISTANCE(mm) y = -0.1971x + 2.406 R² = 0.9722 200 4.00 D(cm) 5.00 6.00 7.00 400 Using your standard curve, calculate the observed size of the two PCR products generated from pOTC and pOTCA and express this base pairs (p) To do this, you will need to measure the migration of the two PCR products on the agerose gel and use the distance migrated to interpolate their logis molendar weght Compare the values you estimate from your standard curve to those you calculated previously in question 12 and provide one way you tright improve the source of the sizes you calculate a standard curve
Expert Answer:
Answer rating: 100% (QA)
To construct a standard curve for the Invitrogen 1 Kb Plus DNA ladder using the migration rates of t... View the full answer

Related Book For 

Fundamentals Of Momentum Heat And Mass Transfer
ISBN: 9781118947463
6th Edition
Authors: James Welty, Gregory L. Rorrer, David G. Foster
Posted Date:
Students also viewed these biology questions
-
Stress (MPa) 500 400 300 200 100 0 Ciled Tensile strength 450 MPa (65,000 psi) MPa 40 200 100 0 10 psi 0.10 30 20 10 0. 0 0.005 Yield strength 250 MPa (36,000 psi) 0.30 E 1 a) Determine the modulus...
-
A ladder 10 ft long rests against a vertical wall. Let be the angle between the top of the ladder and the wall and let be the distance from the bottom of the ladder to the wall. If the bottom of the...
-
The way that PCR amplifies DNA is similar to the doubling in a population of growing bacteria; a single DNA strand is used to synthesize 2 DNA strands, which become 4, then 8, then 16, etc. If a...
-
In the Tokyo subway system, routes are labeled by letters and stops by numbers, such as G-8 or A-3. Stations allowing transfers are sets of stops. Find a Tokyo subway map on the web, develop a simple...
-
On January 1, 2013, Warren Corporation had 1,000,000 shares of common stock outstanding. On March 1, the corporation issued 150,000 new shares to raise additional capital. On July 1, the corporation...
-
Use the info below to answer the following questions: Carnie, Inc purchased bonds with the following assumptions: Par value $ 3 0 0 , 0 0 0 Purchase Price 2 8 2 , 3 3 2 Interest pmts Annual Coupon...
-
The mean room and board expense per year at four-year colleges is \($10,453\). You randomly select 9 four-year colleges. What is the probability that the mean room and board is less than \($10,750?\)...
-
Cycle time efficiency and JIT Walker Brothers Company is considering the installation of a JIT manufacturing system in the hope that it will improve the company??s overall processing cycle...
-
Explain the following statement: Behaviors prime hormones; Hormones prime behavior. Sexual behavior primes aggression; Aggression primes sexual behaviors.
-
John and Ellen Brire are married and file a joint return. They have no dependents. John owns an unincorporated specialty electrical lighting retail store, Brite-On. Brite-On had the following assets...
-
How is the constant evolution of technology beneficial for business? In what ways is it detrimental?
-
Plan corporation acquired 80% of the outstanding voting stock of Sal Corporation on January 1, 2017 for $14,000,000, when Sals stockholders equity consisted of $8,000,000 capital stock and $1,000,000...
-
3. Scenario/Introduction/Background Information: 1909 Drink Sdn Bhd is a newly established company which produces bottled soya bean drink. The company was incorporated on 1" January 2021 and...
-
A beam spanning 7 m face to face between simple supports has a cross-section as shown in Fig. Q-3.3. It is reinforced for flexure with six H32 bars in two layers that continue uninterrupted to the...
-
Kim took out a $15000 loan to start a new business. She can get the loan as an unsecured line of credit at an annual rate of 8.05% compounded daily. If she can afford to pay back $500 per month, how...
-
Problem 3 In the circuit given in the figure I = 2A a) Find the potential difference between the points b and c. b) Find the currents 12 and 13. (The results can be positive or negative) b a 4.0 10.0...
-
You are trying to estimate the cost of equity for a privately owned cruise ship company called the Titanic Cruise Lines. You know that the company's cost of debt is 3% and its debt-to-enterprise...
-
Ex. (17): the vector field F = x i-zj + yz k is defined over the volume of the cuboid given by 0x a,0 y b, 0zc, enclosing the surface S. Evaluate the surface integral ff, F. ds?
-
A flat steel plate of 2.0 m length and 2.0 m width initially contains a very thin coating of light hydrocarbon lubricating oil (species A) used in a manufacturing process. An engineer is considering...
-
At the end of a water pipe of 3-in. diameter is a nozzle that discharges a jet having a diameter of 1 1/2 in into the open atmosphere. The pressure in the pipe is 60 psig (pounds per square inch...
-
The heat loss from a boiler is to be held at a maximum of 900 Btu/h ft 2 of wall area. What thickness of asbestos (k = 0.10 Btu/h ft F) is required if the inner and outer surfaces of the insulation...
-
The trial balance of Shanghai Co. on 31 March 20X7 is given below. The following information is also relevant: 1. Closing inventory is valued at 133m. 2. Electricity accrued is estimated to be 5m. 3....
-
The trial balance for Oslo Co. on 31 July 20X7 is given below. The following additional information is relevant. 1. Closing inventory is valued at 180,000. 2. An allowance for doubtful debts is to be...
-
Find the heat transfer rate \(\mathrm{q}_{\mathrm{w}}\) at \(\mathrm{x}=10 \mathrm{~cm}\) and \(100 \mathrm{~cm}\) for the flat plate given in Problem 7.31. Problem 7.31 A flat plate of \(4...

Study smarter with the SolutionInn App


