A regulatory gene controls the expression of other genesit turns them on and off. Many of these
Question:
A regulatory gene controls the expression of other genes—it turns them on and off. Many of these targets are themselves regulatory genes, instructing genes to turn on or off. Guelzim et al. (2002) determined the number of regulatory genes controlled by a sample of genes in yeast. Their data are listed in the following table.
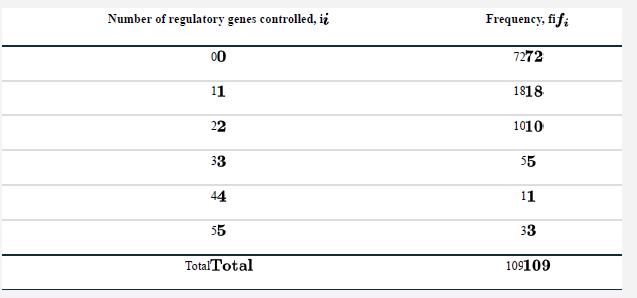
The shape of this frequency distribution suggests that it might be approximated by a probability distribution known as the geometric distribution. Under a geometric distribution, the fraction of genes controlling no regulatory genes (here, i=0 ) is p. The fraction controlling exactly one regulatory gene (i=1) is (1−p)p. The fraction controlling exactly two regulatory genes (i=2) is (1−p)2p, and so on, yielding Pr [i]=(1−p)ip, where i is 0, 1, 2, or more. Maximum likelihood methods can be used to estimate the parameter p, the fraction controlling no regulatory genes, from data. The log-likelihood formula for p is ln[L[ p | data]]=ln[1−p](Σii fi)+n ln[p], where fi is the frequency of observations corresponding to i=0, 1, 2, and so on, and n is the total sample size.
a. Using the data in the table and the preceding formula, calculate the log-likelihood of values of p between 0.1 and 0.9 in increments of 0.01. Use a computer. Draw the relationship between the log-likelihood and p with a log-likelihood curve.
b. What is the maximum likelihood estimate p^, to an accuracy of two decimal places?
c. What is the value of the log-likelihood at the maximum likelihood estimate p^?
d. Using the formula for the geometric distribution, Pr [i]=(1−p)ip, calculate the predicted proportion of genes regulating i=0 to 5 genes based on p^. Plot these values on a histogram with the observed proportions. Based on the result, do the data appear to follow a geometric distribution?
e. Identify one method you might use to test the null hypothesis that a geometric distribution fits the data.
Step by Step Answer:

The Analysis Of Biological Data
ISBN: 9781319226237
3rd Edition
Authors: Michael C. Whitlock, Dolph Schluter





